Detection of virulence, specific genes and antibiotic resistance of isolated Salmonella spp. strains from rabbits infected with salmonellosis
Article information
Abstract
Salmonella spp. are pathogens involved in most salmonellosis in rabbits. This study examined Salmonella disease in rabbits raised in Thua Thien Hue, Vietnam. Two hundred and 56 rectal swabs of rabbits were taken, and a carrier rate of 33.98% was found. In addition, all the isolated Salmonella spp. strains were 100% motile; positive for H2S, catalase, Voges Proskauer, coagulase, citrate, maltose, and dextrose; and negative for indole, methyl red, urease, oxidase, sucrose, and lactose. The Kirby-Bauer method showed that these Salmonella strains were susceptible to doxycycline (93.2%), tetracycline (84.1%), and levofloxacin (65.9%). On the other hand, they were highly resistant to streptomycin (95.5%), ampicillin (93.2%), colistin (40.9%), and gentamicin (34.1%). Furthermore, polymerase chain reaction used to screen for virulence and specific genes of Salmonella strains showed that all Salmonella strains isolated carried InvA, fimA, and Stn.
Introduction
Salmonella is a member of the Enterobacteriaceae family that causes disease in humans, wild animals, and domestic animals [1]. Salmonellosis is one of the most important bacterial infections causing economic losses in livestock because of high mortality and weight loss [2]. Salmonella has been detected in rabbits, posing potential concerns for rabbit farming [3]. Salmonellosis in rabbits has been reported previously [4–8]. The clinical signs of Salmonella infection in rabbits include septicemia, depression, fever, and death, often accompanied by enteritis, diarrhea, metritis, and abortion [9]. Currently, bacterial infections caused by Salmonella spp. in rabbits raised at Thua Thien Hue, Vietnam, remain unknown. Therefore, this study collected 256 rectal swabs of rabbits (both genders aged between 1.5–4 months) from farms and households (16 samples/farm, households) to evaluate the prevalence of Salmonella spp. In addition, biochemical characteristics, common virulence genes, and specific genes were assessed, and their drug resistance characteristics were analyzed to supplement the information as a basis for preventing and controlling Salmonellosis in rabbits in the locality.
Materials and Methods
Sample collection
Two hundred and 56 rectal swabs, including 126 healthy rabbits and 130 rabbits suspected of being infected with Salmonella spp., were collected from 6 households and 10 farms in Thua Thien Hue, Vietnam. Each rectal swab was placed in 5 mL of buffered peptone water (Merck, Germany) and transferred directly to the Laboratory of Immunology and Vaccines, the Institute of Biotechnology, Hue University.
Isolation and characterization of Salmonella spp. from rabbits
The isolation of Salmonella spp. was performed as follows. The samples were incubated overnight at 37°C with shaking. After incubation, 0.1 mL of an overnight culture was added to 10 mL of Rappaport-Vassiliadis, and the mixture was incubated at 42°C for 18 to 24 hours. A loop of each culture was spread onto HiCrome Salmonella Agar (Himedia, India) and incubated at 37°C for 24 hours. The colony morphology was assessed with Gram staining, and biochemical assays were performed: production (H2S, indole, methyl red), catalase, urease activity, Voges Proskauer test, oxidase reaction, coagulase, motility, citrate, fermenting carbohydrates from maltose, sucrose, lactose and dextrose [10]. In addition, presumptive Salmonella spp. colonies were confirmed further by the glass slide agglutination method with multivalent O, H antisera (Denka Seiken Co., Ltd., Japan), according to Grimont and Weill [11].
Antimicrobial sensitivity test
The Kirby-Bauer method [12] was used to obtain antibiograms of all isolated Salmonella strains using 10 antibiotics (Namkhoa, Vietnam): ampicillin 10 µg (Am), amikacin 30 µg (Ak), ceftazidime 30 µg (Cz), chloramphenicol 30 µg (Cl), colistin 10 µg (Co), doxycycline 30 µg (Dx), gentamicin 10 µg (Ge), levofloxacin 5 µg (Lv), streptomycin 10 µg (Sm), and tetracycline 30 µg (Te). The diameters of the inhibition zones on the diffusion disk were measured using the guidelines of the Clinical and Laboratory Standards Institute [13].
Detection of the virulence and specific genes of isolated Salmonella spp.
The genomic DNA of isolated Salmonella spp. was extracted using the TopPURE genomic DNA extraction Kit according to the manufacturer’s guidelines. Azenta Life Sciences synthesized the oligonucleotide primers. Polymerase chain reaction (PCR) was conducted in a total reaction volume of 25 μL (GoTaq Green Master Mix (2X) 12.5µL, DNA template 5 μL, Nuclease-Free Water 5.5 μL, each primer (20 pmol/1 μL)). Table 1 lists the PCR conditions and primer sequences [14,15]. All PCR products were resolved by electrophoresis (1.0% agarose gel in 1X TAE buffer) at 80 V. The gels were stained with GelRed stain and visualized using a UVP UV Transilluminator.
Statistical analysis
The data were processed statistically using the Minitab statistical package ver. 14.0. The results were compared using a chi-squared test, in which p < 0.05 was considered statistically significant.
Results
Prevalence of Salmonella spp. isolated from rabbits
Table 2 lists the prevalence of Salmonella spp. carriers in rabbits raised in Thua Thien Hue. The results revealed an overall carriage rate of 33.98% (87/256). Healthy rabbits are unlikely to be infected with Salmonella spp. The rabbits with diarrhea were infected at a rate of 69.04% (87/126).
Characterization of Salmonella spp. isolates
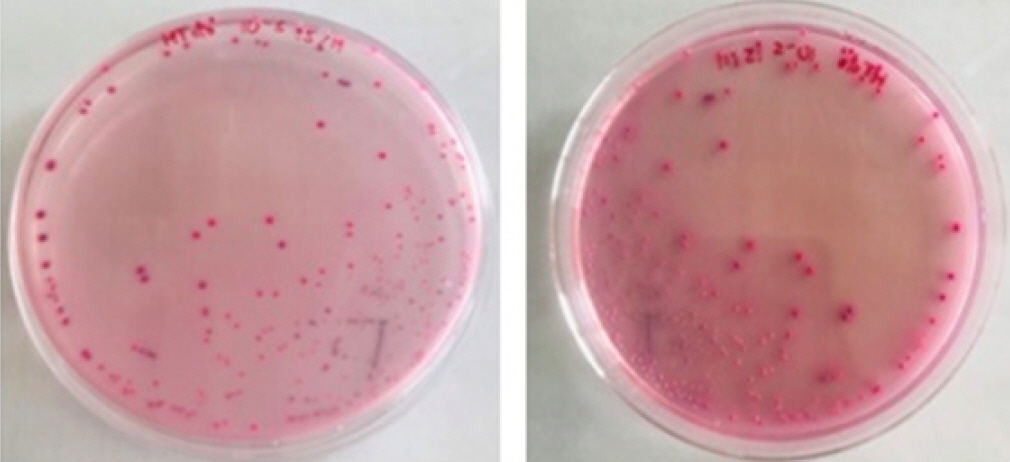
In terms of the morphology, all Salmonella spp. isolated were Gram-negative and rod-shaped. The bacterial colonies appeared pink to red on HiCrome Salmonella Agar (Fig. 1). All strains were motile. They reacted positively to H2S, catalase, Voges Proskauer, coagulase, citrate, maltose, and dextrose and negatively to indole, methyl red, urease, oxidase, sucrose, and lactose. In addition, the slide agglutination test was positive for multivalent O antisera and negative/positive for multivalent H antisera (Table 3).
Determination of antibiotic susceptibility of isolated Salmonella spp.
The isolated Salmonella spp. from rabbits were inhibited by doxycycline (93.2%), tetracycline (84.1%), levofloxacin (65.9%), chloramphenicol/ amikacin (15.9%), colistin (13.6%), ceftazidime (11.4%), and gentamicin (6.8%). On the other hand, they were resistant to streptomycin (95.5%), ampicillin (93.2%), colistin (40.9%), gentamicin (34.1%), amikacin (29.5%), chloramphenicol (11.4%), levofloxacin (4.5%), and ceftazidime (2.3%). Table 4 and Fig. 2 show the antibiotic susceptibility of bacteria.
Detection of virulence and specific genes of Salmonella spp. by PCR
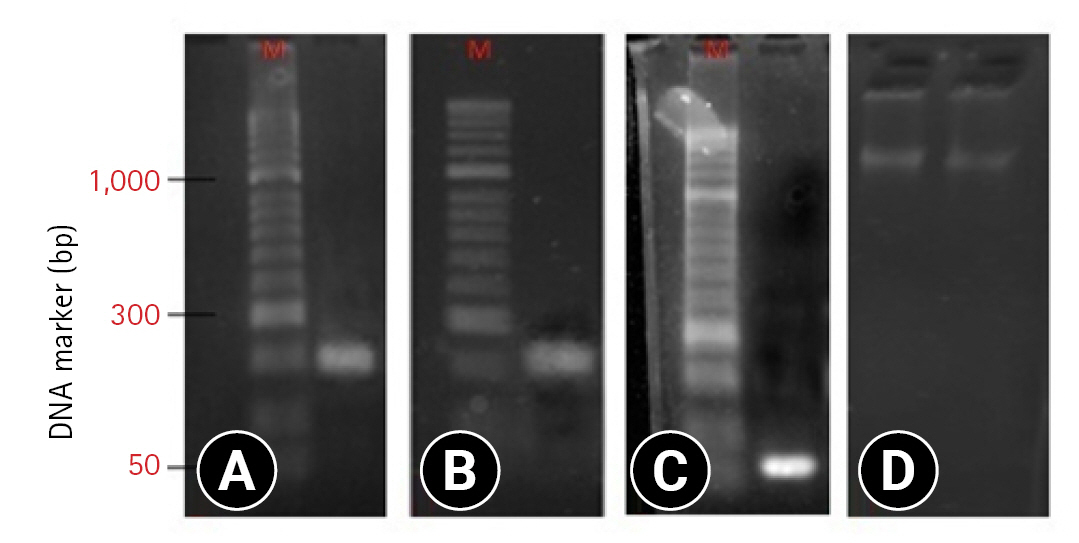
All the Salmonella spp. strains were isolated for DNA extraction, and the presence of invA-specific gene and stn, fimA virulence genes were determined by PCR. The invA, stn, and fimA genes were present in all tested isolates (Fig. 3).
Discussion
Salmonella spp. isolated on HiCrome Salmonella Agar, bacterial colonies appeared pink to red (Fig. 1). This medium is suitable for detecting Salmonella spp. with high specificity [16,17]. Furthermore, all isolated strains could utilize dextrose and maltose but not lactose and sucrose (Table 3), confirming they are Salmonella spp. [18]. The suspected Salmonella spp. strains were motile (Table 3). Salmonella motility is the basic test for detecting motile and non-motile bacteria [19]. Overall, the essential biochemical characteristics of Salmonella spp. in this study were similar to those of other reports [20,21].
Antibiotic resistance in humans and animals is a matter of great concern. Therefore, it is necessary to determine bacterial response to antibiotics so that the correct antibiotics can be employed promptly [22]. This study tested all isolates for susceptibility to 10 commonly used antibiotics in farms/households. Alarmingly, a large percentage of Salmonella spp. isolates displayed strong resistance to streptomycin, ampicillin, and colistin (95.5%, 93.2%, and 40.9%, respectively). On the other hand, the majority were sensitive to doxycycline, tetracycline, and levofloxacin (93.2%, 84.1%, and 65.9%, respectively).
The invA-specific gene has been recognized internationally as the standard for detecting the genus Salmonella [23,24]. InvA gene carries sequences unique to this genus and has been commonly used to detect Salmonella spp. using the PCR method [25]. This gene encodes a protein responsible for the invasion of the host epithelial cells [24,26]. Forty-four conventionally cultured Salmonella spp. strains were selected for DNA extraction and confirmation by PCR. All strains tested carried the invA-specific gene (PCR products of 284 bp in size), confirming that these isolates are Salmonella spp. The results were corroborated by other studies [3,27,28].
The virulence of Salmonella spp. is related to a combination of chromosomal and plasmid factors. There are different genes, of which the Stn is a vital toxin gene encoding the Stn protein that causes gastroenteritis leading to fever, abdominal cramps, nausea, vomiting, and diarrhea. The presence of toxin genes in Salmonella is the cause of salmonellosis and food poisoning diseases in humans [15]. In this study, all Salmonella isolates carried the stn gene with a predicted size of approximately 179 bp. Previously, the stn gene was present in all strains of Salmonella spp. isolated from animals [28,29]. fimA is the gene encoding fimbriae in Salmonella spp. Fimbria-mediated adherence to the intestinal epithelia is a crucial step in enteroaggregative bacterial pathogenesis [30]. The fimA gene was detected in all isolated strains of Salmonella spp., and 85pb DNA fragments were produced, which was similar to that reported by Chaudhary et al. [15] and van Asten and van Dijk [31].
In summary, the Salmonella spp. strains isolated from rabbit rectal swabs were confirmed by bacterial isolation, biochemical reactions, agglutination test, and PCR with primers specific for the InvA gene. Most Salmonella strains carried the genes encoding the enterotoxin stn and the adhesion factor fimA.
Notes
The authors declare no conflict of interest.
Acknowledgements
This study was supported by Hue University through the science and technology project.