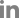Fluoroquinolones (FQs) are a group of antibiotics commonly prescribed for a variety of infections in both human and veterinary medicine. FQs are widely used worldwide because of their bactericidal effects against broad range of bacteria. However, due to the indiscriminate use of FQs, the rise of FQ-resistance in bacteria is regarded as a serious public health concern [
1].
One of the main risk factors for infections caused by antimicrobial-resistant bacteria is contaminated food. Edible offal, which means non-muscular part of the food-producing animalsŌĆÖ carcasses, is a common food product in many countries, but can be easily contaminated by
E. coli present in the intestinal microflora during slaughter and processing [
2]. Therefore, edible offal is a potential source of antimicrobial-resistant
E. coli that can be transferred to humans via the food chain. Although several studies related to FQ-resistant
E. coli in livestock and meat products have been conducted [
3-
5], their significance in edible offal has not been satisfactorily explored. The objective of the current study is to investigate genetic characteristics of FQ-resistant
E. coli isolates from edible offal.
One hundred-eighteen
E. coli isolates were collected from edible offal samples (the heart, liver, stomach or gizzard, small intestine, and large intestine) produced at 8 chicken, 9 pig, and 7 cattle slaughterhouses located in the central and southern regions of Korea from January to October 2017. The method for isolating
E. coli was as follows: 25 g of each sample was incubated in 225 mL of modified EC broth with novobiocin (Merck, Germany) at 37┬░C for 18-24 h. After incubation, 0.1 mL of each broth was inoculated onto MacConkey agar (BD, USA) and incubated at 37┬░C for 18-24 h. Among colonies on the agar, those identified as
malB gene-positive through polymerase chain reaction (PCR) analysis were confirmed as
E. coli [
6]. To find FQ-resistant
E. coli, The
E. coli colonies were streaked onto MacConkey agar containing 4 mg/L ciprofloxacin (Sigma-Aldrich, USA) and incubated at 37┬░C for 18-24 h. Twenty-two FQ-resistant
E. coli isolates (12 isolates from chicken, 8 isolates from pig, and 2 isolates from cattle) were finally collected.
Disk diffusion test was performed to characterize the antimicrobial resistance profiles of FQ-resistant
E. coli isolates according to the Clinical and Laboratory Standards Institute guidelines [
7]. The antimicrobial disks (BD) used were nalidixic acid (NA, 30 ╬╝g), ciprofloxacin (CIP, 5 ╬╝g), ampicillin (AM, 10 ╬╝g), amoxicillin-clavulanate (AMC, 20/10 ╬╝g), cefazolin (CZ, 30 ╬╝g), cefepime (FEP, 30 ╬╝g), cefotaxime (CTX, 30 ╬╝g), cefoxitin (FOX, 30 ╬╝g), cefuroxime (CXM, 30 ╬╝g), ceftazidime (CAZ, 30 ╬╝g), cephalexin (CL, 30 ╬╝g), cephalothin (CF, 30 ╬╝g), chloramphenicol (C, 30 ╬╝g), gentamicin (GM, 10 ╬╝g), imipenem (IPM, 10 ╬╝g), tetracycline (TE, 30 ╬╝g), and trimethoprim-sulfamethoxazole (SXT, 1.25/23.75 ╬╝g). If several isolates from one sample had the same resistance patterns, only one isolate was randomly selected. Multidrug resistance (MDR) was defined as resistance to at least one agent in three or more antimicrobial classes [
8]. Minimum inhibitory concentrations (MICs) of norfloxacin (NOR), CIP, and enrofloxacin (ENR) for FQ-resistant
E. coli isolates were further determined using the agar dilution method.
E. coli ATCC 25922 was used as a control strain.
The FQ-resistant
E. coli isolates were subjected to the PCR method as described previously for detecting the plasmid-mediated quinolone resistance (PMQR) genes (
qnrA,
qnrB,
qnrD,
qnrS,
qepA, and
aac(6ŌĆÖ)-Ib-cr) and the genes causing resistance to ╬▓-lactam antimicrobials (
blaTEM,
blaSHV, and
blaOXA), sulfonamide (
sul1 and
sul2), tetracycline (
tetA and
tetB), chloramphenicol (
catA1 and
cmlA), and aminoglycoside (
aac(6ŌĆÖ)-Ib,
aac(3)-II, and
ant(2ŌĆØ)-I) [
6]. Mutations in the quinolone resistance-determining regions (QRDR) of the
gyrA and
parC genes and other resistance genes were examined by DNA sequencing. The virulence factor genes associated with pathotypes of
E. coli (
eaeA,
escV,
ent,
bfpB,
hly,
stx1,
stx2,
ipaH,
invE,
aggR,
pic,
astA,
elt,
est,
fimH,
papC,
sfa/focDE, and
iucC) were also confirmed by PCR as previously described [
9,
10]. For detecting the 18 major plasmid replicons in Enterobacteriaceae, a PCR-based replicon typing was conducted as described previously [
11].
Pulsed-field gel electrophoresis (PFGE) of the FQ-resistant
E. coli isolates was performed in accordance with CDC PulseNet protocol [
12], using a CHEF-Mapper apparatus (Bio-Rad Laboratories, USA). The dendrogram of PFGE patterns was constructed via Dice coefficients and the unweighted pair group method with arithmetic mean (UPGMA).
The antimicrobial resistance patterns of 22 FQ-resistant
E. coli isolates are shown in
Fig. 1. FQ-resistant
E. coli isolates showed the highest resistance to AM (95.5%) and TE (90.9%), similar to previous studies on the high prevalence of AM and TE resistance from FQ-resistant
E. coli from chicken [
5] and pigs [
3] in Korea. All of the FQ-resistant isolates were also classified as MDR (
Table 1), consistent with the report by Hu et al. [
3] that 91.5% of the FQ-resistant
E. coli isolates showed MDR. Although it is unknown whether there is a direct relationship between FQ-resistance and MDR, this result is not surprising because sales of livestock antimicrobials in Korea have been steadily increasing since 2013, with penicillins and tetracyclines sold more than other antibiotics [
13].
Comparison of MIC values and genetic characteristics related to the FQ-resistance of the isolates are presented in
Fig. 2. The QRDR mutations in FQ target genes, such as
gyrA and
parC, play a significant role in the mechanism of FQ resistance in bacteria [
1]. Similar to previous study in Korea [
3], S83L (18 isolates, 81.8%) and D87N (17 isolates, 77.3%) substations in
gyrA and the S80I substation (20 isolates, 90.9%) in
parC were found to be widespread. Four isolates carried a double mutation in each
gyrA and
parC, and three isolates carried a single mutation in
gyrA and a double mutation in
parC showed the highest MIC ranges (Ōēź 256 ┬Ąg/mL for NOR, 64 to 128 ┬Ąg/mL for CIP, and Ōēź 128 ┬Ąg/mL for ENR). Four isolates which carried a single mutation in both
gyrA and
parC or
gyrA only showed MICs Ōēż 32 ┬Ąg/mL for FQs. In consistent with these results, Moon et al. [
14] also reported that double mutations in
parC led to significantly increased MIC values for FQs. The PMQR genes were detected in 18.2% of FQ-resistant isolates, which is similar to that (15.3%) of pig fecal-derived isolates [
3] and higher than that (5.56%) of chicken isolates [
4] in Korea. Although PMQR genes do not produce high quinolone resistance by themselves, they can facilitate the selection of higher levels of quinolone resistance [
5]. The additional antimicrobial resistance genes
blaTEM-1,
blaOXA-2,
sul1,
sul2,
catA1,
cmlA,
aac(6ŌĆÖ)-Ib,
aac(3)-II,
tetA, and
tetB were also detected, which suggests that FQ-resistant
E. coli may carry resistances against many different antimicrobials as well as deliver FQ-resistance to other bacteria.
A total of five virulence-associated genes (
eaeA,
escV,
astA,
fimH, and
iucC) were found in the isolates tested (
Fig. 2). All 22 FQ-resistant isolates were found to have the
fimH gene in their distribution of virulence genes. The
iuc gene was detected in four isolates, the
astA gene in three isolates, and the
eaeA and
escV genes in one identical isolate. It has been reported that
fimH, the type 1 fimbrial adhesin gene, and
iucC, the aerobactin synthase gene, are common in uropathogenic
E. coli [
10]. In addition, the
eaeA,
escV, and
astA genes contribute to the pathogenicity of diarrheagenic
E. coli [
9].
Also, 17 FQ-resistant isolates were positive for any one of the 18 plasmid replicons (
Fig. 2). Plasmids are small DNA molecules that are distinct from chromosomes and can provide beneficial effects to bacteria such as antibiotic resistance through horizontal gene transfer [
11]. Frep (16 isolates, 94.1%) and FIB (12 isolates, 70.6%), which belong to the IncF group thought to play an important role in the spread of virulence and MDR among Enterobacteriaceae [
15], were more frequent than other replicon types.
XbaI PFGE analysis identified a total of 20 clusters with Ōēź 85% similarity (
Fig. 2). The PFGE patterns of the FQ-resistant isolates revealed generally high genomic diversity (< 50%). However, two isolates obtained from chicken (BC3 and BC6) and pig (QP1 and QP7), respectively, showed the same PFGE patterns. The isolates with identical PFGE patterns were harvested from samples from the same slaughterhouses, suggesting the possibility of cross-contamination of clonal strains during slaughter and processing.
In conclusion, this study provides evidence for the role of edible offal in the dissemination of FQ-resistance in humans via the food chain and shows importance of enhancing hygiene to reduce the cross-contamination in the production of edible offal.











 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Download Citation
Download Citation Print
Print